Explore essential genes
Quick link
If you are looking for bioinformatics knockout SOP, click here to go to the listings. Otherwise, stay and read through the background before looking at the SOP entries.
Objectives
The objective of this page is to present a summary of bioinformatic approaches that can be used to identify whether or not a human gene is likely essential to organismal evolutionary fitness. The page also includes tips on use and interpretation of results from the interactive web server, the Human Essential Genes Interactive Analysis Platform (HEGIAP) (Chen et al 2019).
Note 1: The web server is located via a non-secured hyper-text transfer protocol or http://. This is the older standard, now mostly obsolete. As of March 2026, the folks at HEGIAP have not updated to the secure and encrypted https:// protocol. Therefore your browser is likely to report an error message or strong warning that the site is not secure. Be that as it may, you can view the unsecured site by clicking through the warning message. For example, Chrome browser, select Advanced and then click on “Proceed to [website address] (unsafe)” link.
Background
Essential genes “… are those whose loss of function compromises organism viability or results in profound loss of fitness” (Chen et al 2019 and references therein).
Some genes, that is their products, are more important than others. An extreme example is any gene in which, if its product is absent or malformed, the organism fails to develop or thrive. Now that you have selected a gene of interest (GOI) we begin to ask, do we have evidence that the gene is essential?
Chen et al (2020) summarized shared characteristics of essential genes. Modified from their list, essential genes:
- represent conserved biological processes across species.
- are important for cell development.
- are highly expressed compared to nonessential genes.
- act as ‘hubs’ in protein–protein interaction networks, chromatin structure and epigenetic modification.
A substantial effort of our GWAS essential gene project is to evaluate rates of evolution of our genes of interest, item number 1 from our list of characteristics (see Bioinformatics project: Evolutionary analysis of GWAS candidate genes). Alternatively, we may pursue what is known for our gene of interest with respect to the other characteristics, namely items 2 – 4 from our list.
Inherent to the question is what happens without a functioning copy of the gene, ie, a loss of function mutation, null mutation. Experimentally, we can “knockout” the sequence, preventing the product, then see what happens, from development beginnings throughout life span. Responses range from no change in phenotype across lifespan to stage-specific death. Within that range we can investigate whether how extensive effects to the organism are, from just a handful of phenotypes (monogenic) to “system-wide” effects involving dozens of phenotypes (pleiotropy) or in different ways at different times in the lifespan (antagonistic pleiotropy). While not ethical in human research, model organisms can be used. There are now thousands of results from mouse knockouts on hundreds of genes of interest.
In addition, a substantial number of genes have been evaluated via CRISPR screening. Wang et al (2015) calculated a “CRISPR score,” or CS value. Negative scores suggest the gene is under strong negative selection, that knocking out the gene harms cell survival or growth, and therefore, the gene is likely important for fitness. If the CS score is close to zero (eg, ±0.2), then the gene likely has little to no effect on fitness and the gene is likely non-essential. Conversely, a strongly positive CS score suggests a possible growth advantage if the gene is knocked out. CRISPR score conclusions are context-dependent — cell types tested: for example, a CS score may be strongly negative in a cancer cell line, but zero in a normal cell line; the gene may exhibit synthetic lethality, meaning, the genetic background matters: a gene may only be essential when another gene is mutated or lost.
The HEGIAP web server returns more than the CS score. Highlights include
Epigenetics panels:
- essential genes relatively low methylation of promotor regions, consistent with active transcription (Zhang et al 2015).
- H3K4me3 enriched at promotors near transition start sites or TSS.
- post-transcriptional tri-methylation at the 4th lysine residue of the histone H3 protein.
- H3K27me3 which are often associated with silenced or developmentally regulated genes.
-
- post-transcriptional tri-methylation of the 27th lysine residue on histone H3 protein.
-
- strong 3D chromatin contacts or Hi-C interactions at promoters, consistent with hub-like regulatory regions (Fouziya et al 2024).
Gene structure panels:
- Essential genes tend to be short in length (Kabir et al 2017).
- Few isoforms.
- Conserved exon lengths.
Evolution panels:
- Summarizes results from knockout experiments, including rodents (IMAP and Jackson Labs), yeast, and Drosophila.
- Those genes consistently lethal when knocked out are considered core essential genes, whereas genes which show context-specific lethality are conditionally essential.
- Gene age vs essentiality:
- Evolutionarily ancient (older) genes are more likely essential
- Found across many species
- Involved in core processes (replication, transcription, translation)
- Younger genes more likely specialized, less essential.
- Evolutionarily ancient (older) genes are more likely essential
Chromatin structure panels:
A heatmap of Hi-C promoter interactions vs essentiality. The heatmap shows interaction frequency or contact intensity between genomic regions and is often centered on promoters.
With respect to chromatin structure,
- Essential genes are thought to be enriched in high-contact regions, located in active chromatin compartments, near topologically associating domain (TAD) boundaries or hubs.
- Non-essential genes show fewer interactions and are more isolated or in inactive regions
Note 2: Interpretations derived primarily from Chen et al 2020.
Examples
Examples of essential genes include many so-called housekeeping genes like ACTB. The product, beta-actin is essential to shaping the cell and is involved in cell movement. We can understand why this may be essential, but another way to think about essential is it irreplaceable? Other proteins besides actin are involved in setting cell shape and permitting cell movement. Results of knockout experiments on ACTB. From the HEGIA web server, the CS score of ACTB was -1.648, or the gene is likely essential or strongly fitness-contributing.
Another example, my example gene HIF1A. Review the gene at International Mouse Phenotyping Consortium, results no information, which indicates the experiments have not been tried or are not part of their database. In contrast, Jackson Labs, reports ” Loss of HIF1A results in embryonic lethality.” In contrast, the HEGIA web server, the CS score of HIF1A was 0.379. The interpretation is that HIF1A is likely non-essential under tested conditions.
What to do
As of Spring 2026 semester, I recommend use of the HEGIA web server — see , which provides CS scores and detailed information relevant to characterizing essential genes. Moreover, it combines results from IMPC and Jackson Labs, so as long as the HEGIA server is in operation, there is no benefit to using the other services. However,
Use of IMPC and Jackson Labs to investigate mouse knockout phenotypes
Note 3: The examples were run in 2024. Results may be different today.
International Mouse Phenotyping Consortium
Step 1: Go to the website.
Figure 1. Screenshot of IMPC home page
Note 4: As of 2 February 2024, more than 8000 mouse knockout experiment results are posted at this site. These include results from The Jackson Laboratory, which is a member of the IMPC group — hence the reason I prefer you to use the IMPC site to investigate knockouts involving your GOI.
Step 2: In Search, type your gene of interest name — this example, ACTB — make sure the GENE tab is front.
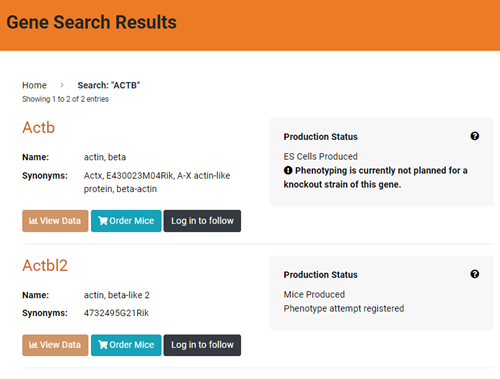
Figure 2. Screenshot of IMPC results for ACTB. Just two strains shown.
Step 3: To view phenotype data, click on the gene link (above “Name”), and record what you find. For example, click on Actb and the page reports:
- Not currently registered for phenotyping at IMPC
- Phenotyping is currently not planned for a knockout strain of this gene.
- which you write into your notebook as your results.
- The related entry Actbl2 instead provided a similar
- Phenotyping is planned for a knockout strain of this gene but data is not currently available.
which you write into your notebook as your results.
End of IMPC SOP.
Step 1: Go to https://mice.jax.org/
Figure 3. Screenshot of JAX homepage.
Step 2: Under FILTERS, click the + next to Strain Attributes and check NULL/KNOCKOUT.
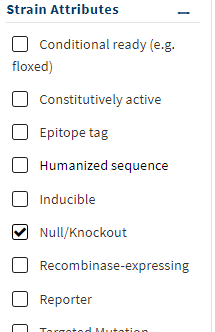
Figure 4. Screenshot of FILTER options. Select NULL/KNOCKOUT.
Note 5: Once you select the filter, the page will update. Check upper left for number of results. As of 2 Feb 2024, 4709 results, i.e., 4709 knockout strains.
Step 3: In Search, type your gene of interest name — this example, ACTB — the site should popup with options that include your keyword. Select just your GOI name, e.g., ACTB.
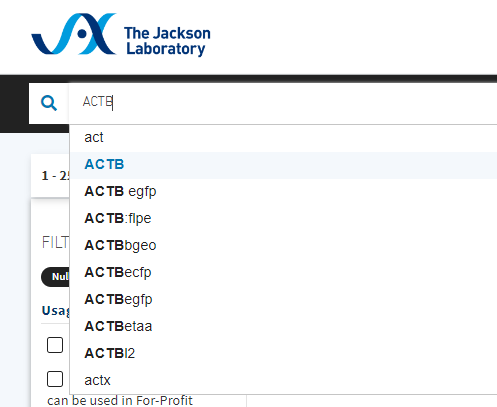
Figure 5. Screenshot JAX search for gene with FILTER = NULL/KNOCKOUT set.
Note 6: Once you enter the search term, the page will update. Check upper left for number of results. As of 2 Feb 2024, 52 results, i.e., 52 knockout strains involving ACTB.
If no results, then report no results! Then, proceed to confirm by checking other databases.
Step 4: In this example, 52 results were presented. For our one credit class, you certainly are not expected to review each of these. (1) Report knockout mice were found (2) Click on any one of the strains listed to review information available about phenotypes and report what you found (you likely need to scroll down the page to view).
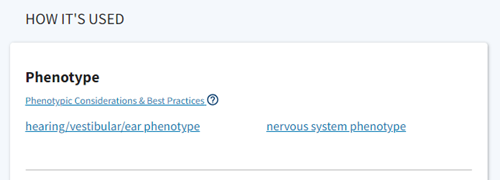
Figure 6. Screenshot of Phenotype results for C57BL/6J-Actbem1Erv/J strain.
Step 5: Explore the phenotypes in more detail. Shown (Fig 7) description after clicking on hearing/vestibular/ear phenotype.
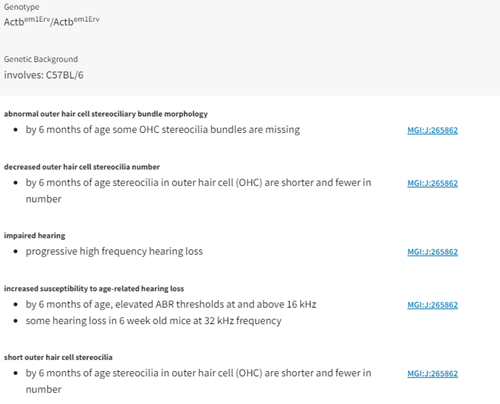
Figure 7. Screenshot hearing/vestibular/ear phenotype descriptions.
And of course, you write results of search findings in your notebook.
End of Jackson Labs SOP.
Use of HEGIAP web server
Note 7: See Note 1 above re: non-secure website browser warning messages.
Step 1. Go to the website.
Figure 8. Screenshot of HEGIAP homepage.
Step 2. Select Start analysis from the menu bar.
Select Analysis single gene from the drop down menu.
Step 3. Enter your gene name in the search bar, then click on Start analysis button.
Figure 9. Screenshot of HEGIAP single gene search page.
Step 4. Review results pages.
In turn, select tabs Epigenetics, Gene structure, Evolution, and Chromatin structure. For brevity, only the top portion of the Epigenetics tab are displayed in Figure 10. Identify the CSA score for your gene:
CS of ACTB: -1.648 top of Epigenetics results pages (Fig 10).
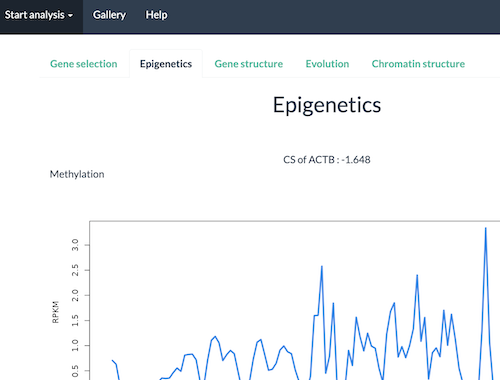
Figure 10. Screenshot of portion of HEGIAP Epigenetics results page for ACTB.
To save results, simply print to pdf and save files to your Working folder.
End of HEGIAP SOP.
Reference s
Chen, H., Zhang, Z., Jiang, S., Li, R., Li, W., Zhao, C., Hong, H., Huang, X., Li, H., & Bo, X. (2020). New insights on human essential genes based on integrated analysis and the construction of the HEGIAP web-based platform. Briefings in Bioinformatics, 21(4), 1397–1410.
Fouziya, S., Krietenstein, N., Mir, U. S., Mieczkowski, J., Khan, M. A., Baba, A., Dar, M. A., Altaf, M., & Wani, A. H. (2024). Genome wide nucleosome landscape shapes 3D chromatin organization. Science Advances, 10(23), eadn2955.
Kabir, M., Barradas, A., Tzotzos, G. T., Hentges, K. E., & Doig, A. J. (2017). Properties of genes essential for mouse development. PLoS ONE, 12(5), e0178273.
Wang, T., Birsoy, K., Hughes, N. W., Krupczak, K. M., Post, Y., Wei, J. J., Lander, E. S., & Sabatini, D. M. (2015). Identification and characterization of essential genes in the human genome. Science, 350(6264), 1096–1101.
Zhang, T., Cooper, S., & Brockdorff, N. (2015). The interplay of histone modifications – writers that read. EMBO Reports, 16(11), 1467–1481.